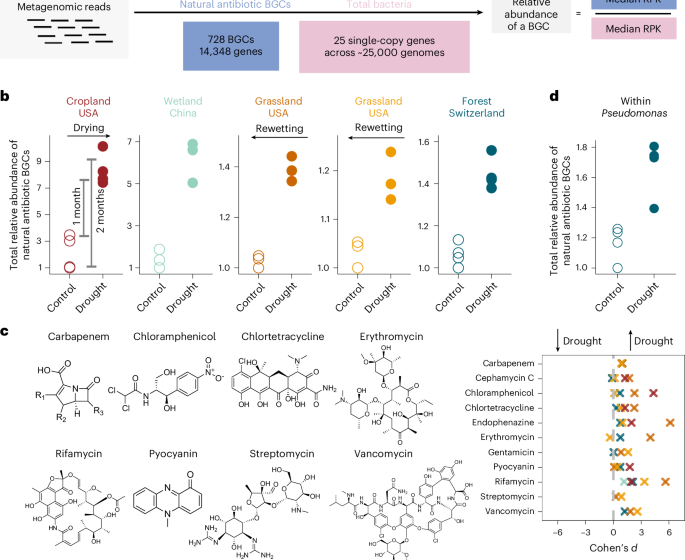
Drought drives elevated antibiotic resistance across soils
Davies, J. & Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 74, 417–433 (2010).
O’Neill, J. Tackling drug-resistant infections globally: final report and recommendations. CABI Digital Library https://www.cabidigitallibrary.org/doi/full/10.5555/20173071720 (2016).
Lewis, K. The science of antibiotic discovery. Cell 181, 29–45 (2020).
Santos-Júnior, C. D. et al. Discovery of antimicrobial peptides in the global microbiome with machine learning. Cell 187, 3761–3778 (2024).
Hover, B. M. et al. Culture-independent discovery of the malacidins as calcium-dependent antibiotics with activity against multidrug-resistant Gram-positive pathogens. Nat. Microbiol. 3, 415–422 (2018).
Lewis, K. Platforms for antibiotic discovery. Nat. Rev. Drug Discov. 12, 371–387 (2013).
Blair, J. M. A., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2014).
Allen, H. K. et al. Call of the wild: antibiotic resistance genes in natural environments. Nat. Rev. Microbiol. 8, 251–259 (2010).
Wright, G. D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 5, 175–186 (2007).
Perry, E. K., Meirelles, L. A. & Newman, D. K. From the soil to the clinic: the impact of microbial secondary metabolites on antibiotic tolerance and resistance. Nat. Rev. Microbiol. 20, 129–142 (2021).
Ault, T. R. On the essentials of drought in a changing climate. Science 368, 256–260 (2020).
Mavrodi, D. V. et al. Accumulation of the antibiotic phenazine-1-carboxylic acid in the rhizosphere of dryland cereals. Appl. Environ. Microbiol. 78, 804–812 (2012).
Mavrodi, D. V. et al. Long-term irrigation affects the dynamics and activity of the wheat rhizosphere microbiome. Front. Plant Sci. 9, 342036 (2018).
Charlop-Powers, Z., Owen, J. G., Reddy, B. V. B., Ternei, M. A. & Brady, S. F. Chemical-biogeographic survey of secondary metabolism in soil. Proc. Natl Acad. Sci. USA 111, 3757–3762 (2014).
Bouskill, N. J. et al. Belowground response to drought in a tropical forest soil. II. Change in microbial function impacts carbon composition. Front. Microbiol. 7, 183812 (2016).
Xu, L. et al. Genome-resolved metagenomics reveals role of iron metabolism in drought-induced rhizosphere microbiome dynamics. Nat. Commun. 12, 3209 (2021).
Santos-Medellín, C., Blazewicz, S. J., Pett-Ridge, J., Firestone, M. K. & Emerson, J. B. Viral but not bacterial community successional patterns reflect extreme turnover shortly after rewetting dry soils. Nat. Ecol. Evol. 7, 1809–1822 (2023).
Hartmann, M. et al. Long-term mitigation of drought changes the functional potential and life-strategies of the forest soil microbiome involved in organic matter decomposition. Front. Microbiol. 14, 1267270 (2023).
Long, Y. et al. Metagenomic analysis revealing the impact of water contents on the composition of soil microbial communities and the distribution of major ecological functional genes in Poyang Lake wetland soil. Microorganisms 12, 2569 (2024).
Medema, M. H. et al. Minimum information about a biosynthetic gene cluster. Nat. Chem. Biol. 11, 625–631 (2015).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Xu, L. et al. Drought delays development of the sorghum root microbiome and enriches for monoderm bacteria. Proc. Natl Acad. Sci. USA 115, E4284–E4293 (2018).
Fitzpatrick, C. R. et al. Assembly and ecological function of the root microbiome across angiosperm plant species. Proc. Natl Acad. Sci. USA 115, E1157–E1165 (2018).
Giacalone, D., Schutt, E. & McRose, D. L. The phospho-ferrozine assay: a tool to study bacterial redox-active metabolites produced at the plant root. Appl. Environ. Microbiol. 91, e02194–24 (2025).
Honeker, L. K. et al. Drought re-routes soil microbial carbon metabolism towards emission of volatile metabolites in an artificial tropical rainforest. Nat. Microbiol. 8, 1480–1494 (2023).
Perry, E. K. & Newman, D. K. Prevalence and correlates of phenazine resistance in culturable bacteria from a dryland wheat field. Appl. Environ. Microbiol. 88, e0232021 (2022).
Dar, D., Thomashow, L. S., Weller, D. M. & Newman, D. K. Global landscape of phenazine biosynthesis and biodegradation reveals species-specific colonization patterns in agricultural soils and crop microbiomes. eLife 9, e59726 (2020).
Meirelles, L. A. & Newman, D. K. Phenazines and toxoflavin act as interspecies modulators of resilience to diverse antibiotics. Mol. Microbiol. 117, 1384–1404 (2022).
Meirelles, L. A., Perry, E. K., Bergkessel, M. & Newman, D. K. Bacterial defenses against a natural antibiotic promote collateral resilience to clinical antibiotics. PLoS Biol. 19, e3001093 (2021).
Russell, A. B., Peterson, S. B. & Mougous, J. D. Type VI secretion system effectors: poisons with a purpose. Nat. Rev. Microbiol. 12, 137–148 (2014).
Tecon, R., Ebrahimi, A., Kleyer, H., Levi, S. E. & Or, D. Cell-to-cell bacterial interactions promoted by drier conditions on soil surfaces. Proc. Natl Acad. Sci. USA 115, 9791–9796 (2018).
Zheng, D. et al. Metagenomics highlights the impact of climate and human activities on antibiotic resistance genes in China’s estuaries. Environ. Pollut. 301, 119015 (2022).
Beaman, B. L. & Beaman, L. Nocardia species: host–parasite relationships. Clin. Microbiol. Rev. 7, 213–264 (1994).
Kirby, R. et al. Draft genome sequence of the human pathogen streptomyces somaliensis, a significant cause of actinomycetoma. J. Bacteriol. 194, 3544–3545 (2012).
Smith, I. Mycobacterium tuberculosis pathogenesis and molecular determinants of virulence. Clin. Microbiol. Rev. 16, 463–496 (2003).
Scollard, D. M. et al. The continuing challenges of leprosy. Clin. Microbiol. Rev. 19, 338–381 (2006).
Forsberg, K. J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Jiang, X. et al. Dissemination of antibiotic resistance genes from antibiotic producers to pathogens. Nat. Commun. 8, 15784 (2017).
Mahmud, B. et al. Longitudinal dynamics of farmer and livestock nasal and faecal microbiomes and resistomes. Nat. Microbiol. 9, 1007–1020 (2024).
Mhuireach, G., Van Den Wymelenberg, K. G. & Langellotto, G. A. Garden soil bacteria transiently colonize gardeners’ skin after direct soil contact. Urban Agric. Region. Food Syst. 8, e20035 (2023).
Hartmann, E. M. et al. Antimicrobial chemicals are associated with elevated antibiotic resistance genes in the indoor dust microbiome. Environ. Sci. Technol. 50, 9807–9815 (2016).
Zhou, Z. et al. Association between particulate matter (PM)2·5 air pollution and clinical antibiotic resistance: a global analysis. Lancet Planet. Health 7, e649–e659 (2023).
De Martonne, E. Une nouvelle function climatologique: L’indice d’aridite. Meteorologie 2, 449–459 (1926).
O’Neill, B. C. et al. The roads ahead: narratives for shared socioeconomic pathways describing world futures in the 21st century. Global Environ. Change 42, 169–180 (2017).
Kelley, M. et al. GISS-E2.1: configurations and climatology. J. Adv. Model. Earth Syst. 12, e2019MS002025 (2020).
Gutjahr, O. et al. Max Planck Institute Earth System Model (MPI-ESM1.2) for the High-Resolution Model Intercomparison Project (HighResMIP). Geosci. Model Dev. 12, 3241–3281 (2019).
Andrews, M. B. et al. Historical simulations With HadGEM3-GC3.1 for CMIP6. J. Adv. Model. Earth Syst. 12, e2019MS001995 (2020).
Li, X. J. et al. Uncertainties in global permafrost area extent estimates from different methods. Adv. Clim. Chang. Res. 16, 312–323 (2025).
Falghoush, A., Beyenal, H., Besser, T. E., Omsland, A. & Call, D. R. Osmotic compounds enhance antibiotic efficacy against Acinetobacter baumannii biofilm communities. Appl. Environ. Microbiol. 83, e01297–17 (2017).
Barraud, N., Buson, A., Jarolimek, W. & Rice, S. A. Mannitol enhances antibiotic sensitivity of persister bacteria in Pseudomonas aeruginosa biofilms. PLoS ONE 8, e84220 (2013).
Al-Nabulsi, A. A. et al. Effects of osmotic pressure, acid, or cold stresses on antibiotic susceptibility of Listeria monocytogenes. Food Microbiol. 46, 154–160 (2015).
Hood, M. I., Jacobs, A. C., Sayood, K., Dunman, P. M. & Skaar, E. P. Acinetobacter baumannii increases tolerance to antibiotics in response to monovalent cations. Antimicrob. Agents Chemother. 54, 1029–1041 (2010).
Windham, I. H. & Merrell, D. S. Interplay between amoxicillin resistance and osmotic stress in Helicobacter pylori. J. Bacteriol. 204, e0004522 (2022).
Xu, X. et al. Adsorption/desorption and degradation of doxycycline in three agricultural soils. Ecotoxicol. Environ. Saf. 224, 112675 (2021).
Huang, S. et al. Dry-to-wet fluctuation of moisture contents enhanced the mineralization of chloramphenicol antibiotic. Water Res. 240, 120103 (2023).
Hernando-Amado, S., Coque, T. M., Baquero, F. & Martínez, J. L. Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nat. Microbiol. 4, 1432–1442 (2019).
Zambrano, M. M. Interplay between antimicrobial resistance and global environmental change. Annu. Rev. Genet. 57, 275–296 (2023).
Alcock, B. P. et al. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 51, D690–D699 (2023).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2014).
Zhang, J. et al. SecReT6 update: a comprehensive resource of bacterial type VI secretion systems. Sci. China Life Sci. 66, 626–634 (2022).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
visNetwork, an R package for interactive network visualization. GitHub http://datastorm-open.github.io/visNetwork/ (2022).
GNI per capita, Atlas method (current US$). World Bank Group http://data.worldbank.org/indicator/NY.GNP.PCAP.CD (2025).
Thrasher, B. et al. NASA global daily downscaled projections. CMIP6. Sci. Data 9, 262 (2022).
Thrasher B., Wang W., Michaelis A. & Nemani R. NEX-GDDP-CMIP6. NASA Center for Climate Simulations https://doi.org/10.7917/OFSG3345 (2021).
Eyring, V. et al. Overview of the Coupled Model Intercomparison Project Phase 6 (CMIP6) experimental design and organization. Geosci. Model Dev. 9, 1937–1958 (2016).
Hoyer, S. & Hamman, J. J. xarray: N-D labeled arrays and datasets in Python. J. Open Res. Softw. 5, 10 (2017).
Cartopy—a cartographic python library with matplotlib support. GitHub http://github.com/SciTools/cartopy (2026).
Ben-Shachar, M. S., Lüdecke, D. & Makowski, D. Effectsize: estimation of effect size indices and standardized parameters. J. Open Source Softw. 5, 2815 (2020).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2017).
Cohen, J. Statistical Power Analysis for the Behavioral Sciences 2nd edn. https://doi.org/10.4324/9780203771587 (Routledge, 1988).
First Appeared on
Source link






