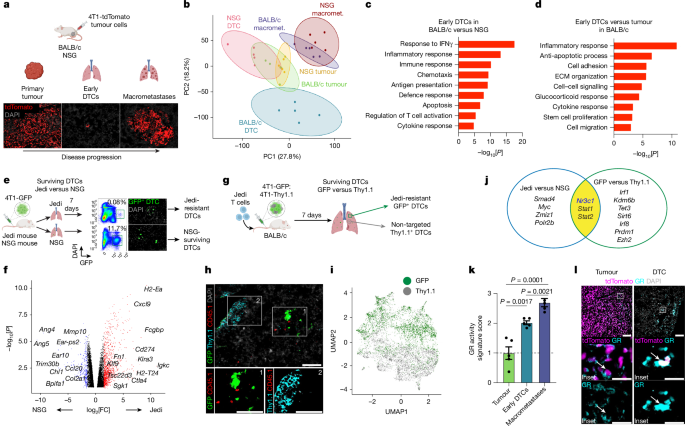
A glucocorticoid–FAS axis controls immune evasion during metastatic seeding
Lambert, A. W., Zhang, Y. & Weinberg, R. A. Cell-intrinsic and microenvironmental determinants of metastatic colonization. Nat. Cell Biol. 26, 687–697 (2024).
Garrido-Castro, A. C., Lin, N. U. & Polyak, K. Insights into molecular classifications of triple-negative breast cancer: improving patient selection for treatment. Cancer Discov. 9, 176–198 (2019).
Pogoda, K., Niwińska, A., Murawska, M. & Pieńkowski, T. Analysis of pattern, time and risk factors influencing recurrence in triple-negative breast cancer patients. Med. Oncol. 30, 388 (2013).
Harper, K. L. et al. Mechanism of early dissemination and metastasis in Her2+ mammary cancer. Nature 540, 588–592 (2016).
Hosseini, H. et al. Early dissemination seeds metastasis in breast cancer. Nature 540, 552–558 (2016).
Hüsemann, Y. et al. Systemic spread is an early step in breast cancer. Cancer Cell 13, 58–68 (2008).
Goddard, E. T. et al. Immune evasion of dormant disseminated tumor cells is due to their scarcity and can be overcome by T cell immunotherapies. Cancer Cell 42, 119–134 (2024).
Michaels, E., Chen, N. & Nanda, R. The role of immunotherapy in triple-negative breast cancer (TNBC). Clin. Breast Cancer 24, 263–270 (2024).
Kroemer, G., Senovilla, L., Galluzzi, L., Andre, F. & Zitvogel, L. Natural and therapy-induced immunosurveillance in breast cancer. Nat. Med. 21, 1128–1138 (2015).
Baldominos, P. et al. Quiescent cancer cells resist T cell attack by forming an immunosuppressive niche. Cell 185, 1694–1708 (2022).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Malladi, S. et al. Metastatic latency and immune evasion through autocrine inhibition of WNT. Cell 165, 45–60 (2016).
Baumann, Z., Auf der Maur, P. & Bentires-Alj, M. Feed-forward loops between metastatic cancer cells and their microenvironment-the stage of escalation. EMBO Mol. Med. 14, e14283 (2022).
Massague, J. & Ganesh, K. Metastasis-initiating cells and ecosystems. Cancer Discov. 11, 971–994 (2021).
Hu, J. et al. STING inhibits the reactivation of dormant metastasis in lung adenocarcinoma. Nature 616, 806–813 (2023).
Agudo, J. et al. GFP-specific CD8 T cells enable targeted cell depletion and visualization of T-cell interactions. Nat. Biotechnol. 33, 1287–1292 (2015).
Falcinelli, M. et al. The role of psychologic stress in cancer initiation: clinical relevance and potential molecular mechanisms. Cancer Res. 81, 5131–5140 (2021).
Abduljabbar, R. et al. Clinical and biological significance of glucocorticoid receptor (GR) expression in breast cancer. Breast Cancer Res. Treat. 150, 335–346 (2015).
Chen, Z. et al. Ligand-dependent genomic function of glucocorticoid receptor in triple-negative breast cancer. Nat. Commun. 6, 8323 (2015).
Pan, D., Kocherginsky, M. & Conzen, S. D. Activation of the glucocorticoid receptor is associated with poor prognosis in estrogen receptor-negative breast cancer. Cancer Res. 71, 6360–6370 (2011).
West, D. C. et al. Discovery of a glucocorticoid receptor (GR) activity signature using selective GR antagonism in ER-negative breast cancer. Clin. Cancer Res. 24, 3433–3446 (2018).
Obradovic, M. M. S. et al. Glucocorticoids promote breast cancer metastasis. Nature 567, 540–544 (2019).
Flaherty, R. L. et al. Stress hormone-mediated acceleration of breast cancer metastasis is halted by inhibition of nitric oxide synthase. Cancer Lett. 459, 59–71 (2019).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Pommier, A. et al. Unresolved endoplasmic reticulum stress engenders immune-resistant, latent pancreatic cancer metastases. Science 360, eaao4908 (2018).
Qin, Q. et al. Lisa: inferring transcriptional regulators through integrative modeling of public chromatin accessibility and ChIP-seq data. Genome Biol. 21, 32 (2020).
Garcia-Recio, S. et al. Multiomics in primary and metastatic breast tumors from the AURORA US network finds microenvironment and epigenetic drivers of metastasis. Nat. Cancer 4, 128–147 (2023).
Mayer, E. L. et al. TBCRC 030: a phase II study of preoperative cisplatin versus paclitaxel in triple-negative breast cancer: evaluating the homologous recombination deficiency (HRD) biomarker. Ann. Oncol. 31, 1518–1525 (2020).
Parsons, H. A. et al. Circulating tumor DNA association with residual cancer burden after neoadjuvant chemotherapy in triple-negative breast cancer in TBCRC 030. Ann. Oncol. 34, 899–906 (2023).
Ombrato, L. et al. Metastatic-niche labelling reveals parenchymal cells with stem features. Nature 572, 603–608 (2019).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Gerber, A. N., Newton, R. & Sasse, S. K. Repression of transcription by the glucocorticoid receptor: a parsimonious model for the genomics era. J. Biol. Chem. 296, 100687 (2021).
Hudson, W. H. et al. Cryptic glucocorticoid receptor-binding sites pervade genomic NF-κB response elements. Nat. Commun. 9, 1337 (2018).
Liu, F. et al. NF-κB directly regulates Fas transcription to modulate Fas-mediated apoptosis and tumor suppression. J. Biol. Chem. 287, 25530–25540 (2012).
Zheng, Y. et al. NF-kappa B RelA (p65) is essential for TNF-alpha-induced fas expression but dispensable for both TCR-induced expression and activation-induced cell death. J. Immunol. 166, 4949–4957 (2001).
Singh, R., Letai, A. & Sarosiek, K. Regulation of apoptosis in health and disease: the balancing act of BCL-2 family proteins. Nat. Rev. Mol. Cell Biol. 20, 175–193 (2019).
Kaufmann, T., Strasser, A. & Jost, P. J. Fas death receptor signalling: roles of Bid and XIAP. Cell Death Differ. 19, 42–50 (2012).
Sarosiek, K. A. et al. BID preferentially activates BAK while BIM preferentially activates BAX, affecting chemotherapy response. Mol. Cell 51, 751–765 (2013).
Hopkins, S. & Fleseriu, M. in Cushing’s Disease (Laws, E. R.) 103–123 (Academic Press, 2017).
Yang, H. et al. Stress-glucocorticoid-TSC22D3 axis compromises therapy-induced antitumor immunity. Nat. Med. 25, 1428–1441 (2019).
Zhou, P. et al. In vivo discovery of immunotherapy targets in the tumour microenvironment. Nature 506, 52–57 (2014).
Dupaul-Chicoine, J. et al. The Nlrp3 inflammasome suppresses colorectal cancer metastatic growth in the liver by promoting natural killer cell tumoricidal activity. Immunity 43, 751–763 (2015).
Valiente, M. et al. Serpins promote cancer cell survival and vascular co-option in brain metastasis. Cell 156, 1002–1016 (2014).
van der Pompe, G., Antoni, M. H. & Heijnen, C. J. Elevated basal cortisol levels and attenuated ACTH and cortisol responses to a behavioral challenge in women with metastatic breast cancer. Psychoneuroendocrinology 21, 361–374 (1996).
Acharya, N. et al. Endogenous glucocorticoid signaling regulates CD8+ T cell differentiation and development of dysfunction in the tumor microenvironment. Immunity 53, 658–671 (2020).
He, X. Y. et al. Chronic stress increases metastasis via neutrophil-mediated changes to the microenvironment. Cancer Cell 42, 474–486 (2024).
Budiu, R. A. et al. Restraint and social isolation stressors differentially regulate adaptive immunity and tumor angiogenesis in a breast cancer mouse model. Cancer Clin. Oncol. 6, 12–24 (2017).
Freed-Pastor, W. A. et al. The CD155/TIGIT axis promotes and maintains immune evasion in neoantigen-expressing pancreatic cancer. Cancer Cell 39, 1342–1360 (2021).
Brown, B. D., Venneri, M. A., Zingale, A., Sergi Sergi, L. & Naldini, L. Endogenous microRNA regulation suppresses transgene expression in hematopoietic lineages and enables stable gene transfer. Nat. Med. 12, 585–591 (2006).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Patel, S. G. et al. Cell-penetrating peptide sequence and modification dependent uptake and subcellular distribution of green florescent protein in different cell lines. Sci. Rep. 9, 6298 (2019).
Kim, W. J., Kim, G. R., Cho, H. J. & Choi, J. M. The cysteine-containing cell-penetrating peptide AP enables efficient macromolecule delivery to T cells and controls autoimmune encephalomyelitis. pharmaceutics. Pharmaceutics 13, 1134 (2021).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
McCarthy, D. J., Campbell, K. R., Lun, A. T. L. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Germain, P. L., Lun, A., Garcia Meixide, C., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res 10, 979 (2021).
Hsu, L. L. & Culhane, A. C. Correspondence analysis for dimension reduction, batch integration, and visualization of single-cell RNA-seq data. Sci. Rep. 13, 1197 (2023).
Lun, A. T., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res 5, 2122 (2016).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Langmead, B. & Nellore, A. Cloud computing for genomic data analysis and collaboration. Nat. Rev. Genet. 19, 208–219 (2018).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Zhang, Y. et al. Model-based Analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Certo, M. et al. Mitochondria primed by death signals determine cellular addiction to antiapoptotic BCL-2 family members. Cancer Cell 9, 351–365 (2006).
Fraser, C., Ryan, J. & Sarosiek, K. BH3 profiling: a functional assay to measure apoptotic priming and dependencies. Methods Mol. Biol. 1877, 61–76 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Posta, M. & Győrffy, B. Pathway-level mutational signatures predict breast cancer outcomes and reveal therapeutic targets. Br. J. Pharmacol. 182, 5734–5747 (2025).
Tang, W. et al. Integrated proteotranscriptomics of breast cancer reveals globally increased protein-mRNA concordance associated with subtypes and survival. Genome Med. 10, 94 (2018).
Gyorffy, B. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput. Struct. Biotechnol. J. 19, 4101–4109 (2021).
First Appeared on
Source link






